Introduction to machine learning for bioinformatics
Machine learning is a subset of the broader field of artificial intelligence (AI). It enables systems to independently learn from data and execute tasks that they are not explicitly programmed to handle. Its goal is to give machines the ability to perform tasks that require human intelligence, such as diagnosing, planning, and predicting.
There are two main types of machine learning:
-
Supervised learning relies on labeled datasets to teach algorithms an existing classification system and how to make predictions based on it. This ML type is used to train decision trees and neural networks.
-
Unsupervised learning doesn’t use labels. Instead, algorithms try to uncover data patterns on their own. In other words, they learn things that we can’t teach them directly. This is comparable to how the human brain works.
It’s also possible to combine labeled and unlabeled data during training, which will result in semi-supervised learning. This ML type can be useful when you don’t have enough high-quality labeled data for a supervised learning approach, but you still want to use it to direct the learning process.

What are the most popular machine learning techniques used in bioinformatics?
Some of these algorithms fall strictly under the supervised/unsupervised learning categories, and some can be used with both methods.
Natural language processing
Natural language processing (NLP) is a set of techniques that can understand unstructured human language.
NLP can search through volumes of biology research, aggregate information on a given topic from various sources, and translate research findings from one language to another. In addition to mining research papers, NLP solutions can parse relevant biomedical databases.
NLP can benefit the bioinformatics field in the following ways:
-
Interpreting genetic variants
-
Analyzing DNA expression arrays
-
Annotating protein functions
-
Looking for new drug targets
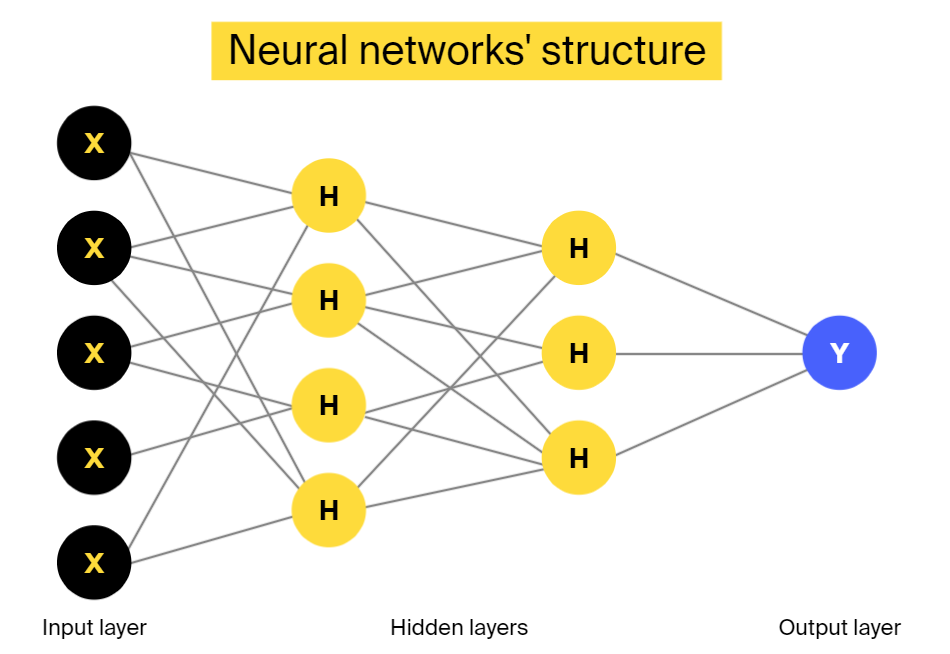
Neural networks
This is a multi-layered structure consisting of nodes/neurons as its building blocks. Neurons in adjacent layers are connected to each other via links, but neurons of the same layer are not interlinked. The input layer neurons receive information, process it, and pass it along as an input to the next layer. And this process continues until the processed information reaches the output layer.
The most basic neural network is called perceptron. It consists of one neuron that acts as a classifier. This neuron receives an input and places it in one of two classes using a linear discrimination function. In larger neural networks, there is no limit on the number of layers or the number of nodes in one layer.
-
Classifying gene expression profiles
-
Predicting protein structure
-
Sequencing DNA
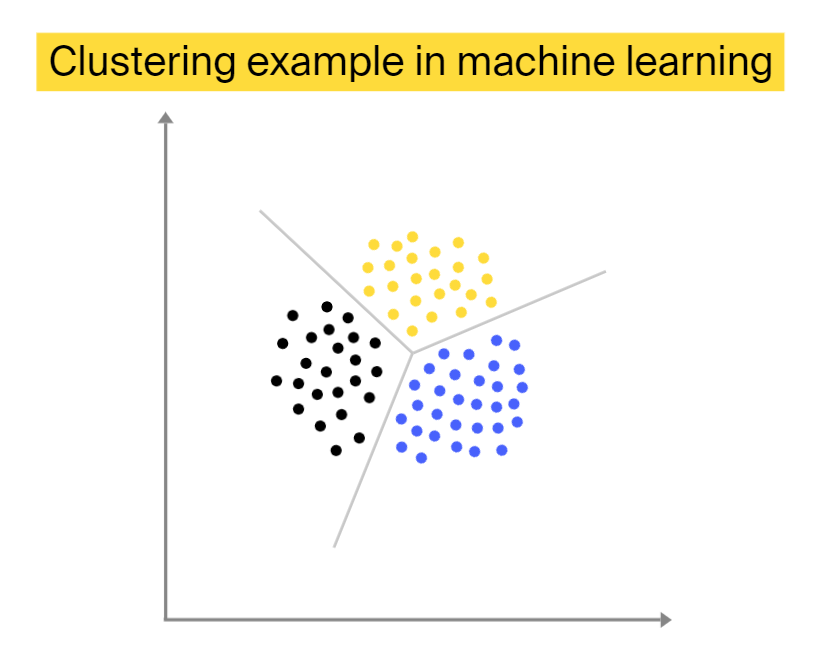
Clustering
Unsupervised clustering is the process of organizing elements into various groups based on the supplied definition of similarity. As a result of such classification, the elements positioned in one cluster closely relate to one another, and differ from elements in other clusters.
Unlike with supervised classification, in clustering, we don’t know in advance how many clusters will be formed. One famous example of this machine learning approach in bioinformatics is microarray-based expression profiling of genes, where genes with similar expression levels are positioned in one cluster.
Dimensionality reduction
In machine learning classification problems, classifications are performed based on factors/features. Sometimes there are too many factors that affect the final result, making the dataset difficult to visualize and manipulate. Dimensionality reduction algorithms can minimize the number of features, making the dataset more manageable. For instance, a climate classification problem might have humidity and rainfall among its features. These two can be collapsed into one factor for the sake of simplicity as they are both closely related.
Dimensionality reduction has two main components:
-
Feature selection. Chooses a subset of variables to represent the entire model by embedding, filtering, or wrapping features.
-
Feature extraction. Reduces the number of dimensions in a dataset. For instance, a 3D space can be broken into two 2D spaces.
This type of algorithms is used to compress large datasets for the sake of reducing computational time and storage requirements. It can also eliminate redundant features present in the data.
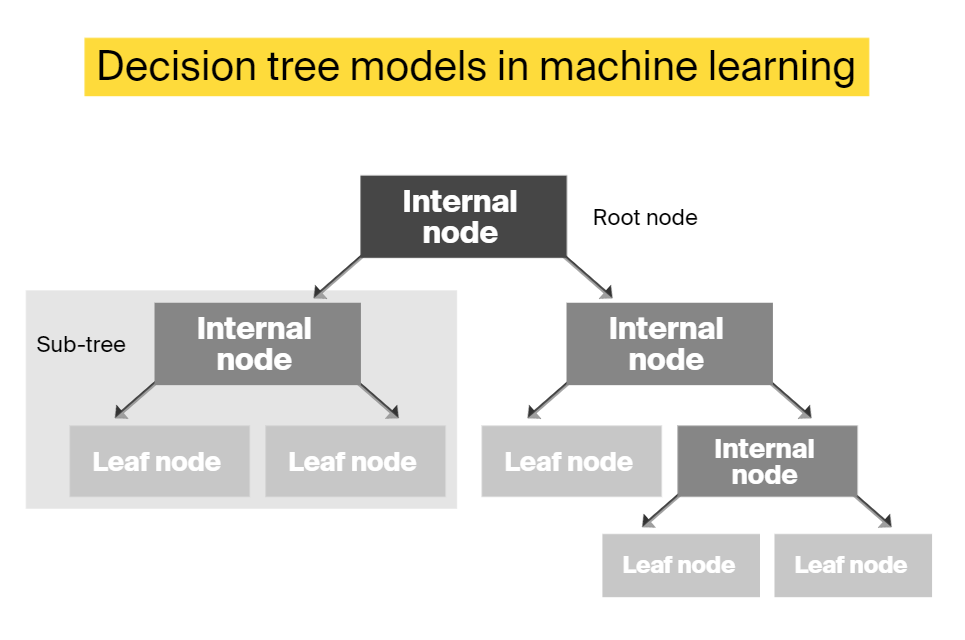
Decision tree classifiers
This is one of the most popular classical supervised learning classifiers. These algorithms apply a recursive approach to build a flowchart-like tree model, where each node represents a test on a feature. First, the algorithm determines the top node — the root — and then builds the tree recursively considering one parameter at a time. The final node in each sequence is called “the leaf node.” It represents the final classification and holds the class label.
Decision tree models demand high computational power during training, but afterwards they can perform classifications without extensive computing. The main advantage these classifiers bring to the bioinformatics field is that they generate understandable rules and explainable results.
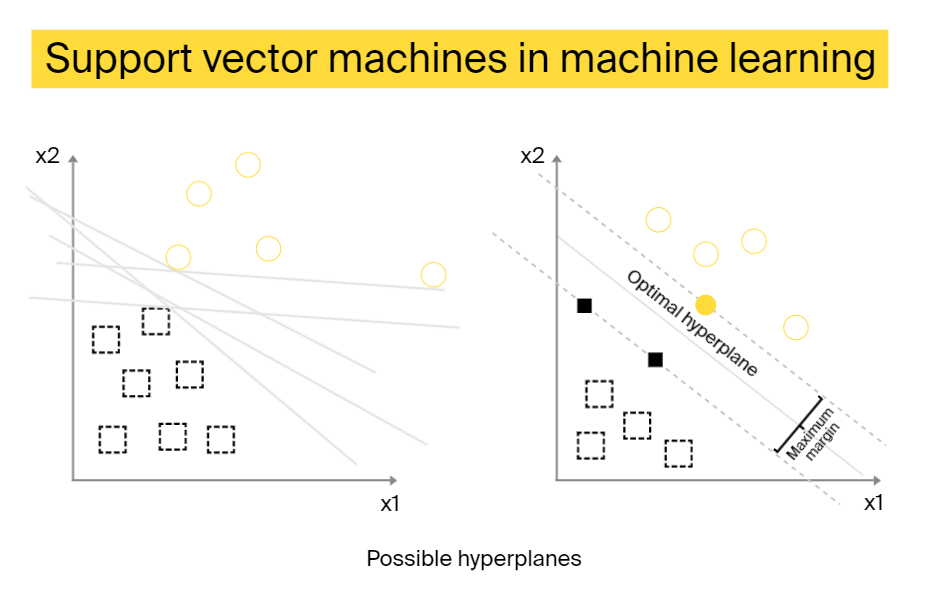
Support vector machine
This is a supervised ML model that can solve two-group classification problems. To classify data points, these algorithms look for an optimal hyperplane that divides the data separating it into two classes with the maximum distance between data points.
The points located on either sides of the hyperplane belong to different classes. The hyperplane’s dimension depends on the number of features. In the case of two features, the decision boundary is a line, with three features, it’s a 2D plate. This characteristic makes it hard to use SVM for classifications with more than three features.
This approach is useful in computational identification of functional RNA genes. It can select the optimal set of genes for cancer detection based on their expression data.
Top 5 applications of machine learning in bioinformatics
After giving a brief introduction to machine learning and highlighting the most commonly used ML algorithms, let’s see how they can be deployed in the bioinformatics field.
If any of these use cases catches your attention, turn to AI software consulting professionals to implement a customized solution for your business.
1. Facilitating gene editing experiments
Gene editing refers to manipulations on an organism’s genetic composition by deleting, inserting, and replacing a part of its DNA sequence. This process typically relies on the CRISPR technique, which is rather effective. But there is still much improvement to be desired in the area of selecting the right DNA sequence for manipulation, and this is where ML can help. Using machine learning for bioinformatics, researchers can enhance the design of gene editing experiments and predict their outcomes.
A research team employed ML algorithms to discover the most optimal combinational variants of amino-acid residues that allow genome-editing protein Cas9 to bind with the target DNA. Due to the large number of these variants, such an experiment would have been too large, but using an ML-driven engineering approach reduced the screening burden by around 95%.
2. Identifying protein structure
Proteomics is a study of proteins, their interactions, composition, and their role in the human body. This field involves heavy biological datasets and is computationally expensive. Therefore, technologies like machine learning in bioinformatics are essential here.
One of the most successful applications in this field is using convolutional neural networks to position proteins’ amino acids into three classes — sheet, helix, and coil. Neural networks can achieve an accuracy of 84% with the theoretical limit being 88%–90%.
Another usage of ML in proteomics is protein model scoring, a task essential to predict protein structure. In their machine learning approach to bioinformatics, researchers from the Fayetteville State University deployed ML to improve protein model scoring. They divided protein models under question into groups and used an ML interpreter to decide on the feature vector to evaluate models belonging to each group. These feature vectors were used later to further improve the ML algorithms while training them on each group separately.
3. Spotting genes associated with diseases
Researchers increasingly use machine learning in bioinformatics to identify genes that are likely to be involved in particular diseases. This is achieved by analyzing gene expression microarrays and RNA sequencing.
Particularly, gene identification gains traction in cancer-related studies to identify genes that are likely to contribute to cancer, as well as classify tumors by analyzing them on a molecular level.
For instance, a group of scientists at the University of Washington used several machine learning in bioinformatics algorithms, including decision tree, support vector machine, and neural networks to test their ability to predict and classify cancer types. Researchers deployed RNA sequencing data from The Cancer Genome Atlas project, and discovered that linear support vector machine was the most precise, hitting the 95.8% accuracy in cancer classification.
In another example, researchers used ML to classify breast cancer types based on gene expression data. This team also relied on the Cancer Genome Atlas project’s data. The researchers classified the samples into triple negative breast cancer — one of the most lethal breast cancers — and non-triple negative. And once again, the support vector machine classifier delivered the best results.
Speaking of non-cancerous diseases, researchers at the University of Pennsylvania relied on machine learning to identify genes that would be a suitable target for coronary artery disease (CAD) drugs. The team used the ML-powered Tree-based Pipeline Optimization Tool (TPOT) to pinpoint a combination of single nucleotide polymorphisms (SNPs) related to CAD. They analyzed the genomic data from the UK Biobank and uncovered 28 relevant SNPs. The relation between the SNPs on top of this list and CAD was previously mentioned in the literature, and this research gave a practical validation.
4. Traversing the knowledge base in search of meaningful patterns
Advanced sequencing technology doubles genomic databases each 2.5 years, and researchers are looking for a way to extract useful insights from this accumulated knowledge. Machine learning in bioinformatics can sift through biomedical publications and reports to identify different genes and proteins and search for their functionality. It can also aid in annotating protein databases and complement them with the information it retrieves from the literature.
One example comes from a group of researchers who deployed bioinformatics and machine learning in literature mining to facilitate protein model scoring. Structural modeling of protein-protein dockings typically results in several models that are further scored based on structural constraints. The team used ML algorithms to traverse PubMed papers on protein-protein interactions, searching for residues that could help generate these constraints for model scoring. And to make sure that the constraints are relevant, scientists explored the ability of different machine learning algorithms to check all discovered residues for relevancy.
This research revealed that both computationally expensive neural networks and less resource demanding support vector machine achieved very similar results.
5. Repurposing drugs
Drug repurposing, or reprofiling, is a technique scientists use to discover new applications of existing drugs that they were not intended for. Researchers adopt AI in bioinformatics to perform drug analysis on relevant databases, such as BindingDB and DrugBank. There are three major directions for drug repurposing:
-
Drug-target interaction looks into the drug’s ability to bind directly to the target protein
-
Drug-drug interaction investigates how medications act when they are taken in combinations
-
Protein-protein interaction looks into the surface of interacting intracellular proteins, and attempts to discover hotspots and allosteric sites.
Researchers from the China University of Petroleum and the Shandong University developed a deep neural network algorithm and used it on the DrugBank database. They wanted to study drug-target interactions between drug molecules and the mitochondrial fusion protein 2 (MFN2), which is one of the main proteins that can possibly cause Alzheimer’s disease. The study identifies 15 drug molecules with binding potential. Upon further investigation, it appeared that 11 of them can successfully dock with MFN2. And five of them have medium to strong binding force.
Challenges presented by machine learning in bioinformatics
Machine learning in bioinformatics differs from ML in other sectors due to the four factors below, which also constitute the main challenges of applying ML to this field.
-
Bioinformatics AI is expensive. For the algorithm to perform properly, you need to acquire a large training dataset. However, it’s rather costly to obtain 10,000 chest scans, or any other type of medical data for that matter.
-
Difficulties associated with the training datasets. In other fields, if you don’t have enough training data, you can generate synthetic data to expand your dataset. However, this trick might not be appropriate when it comes to human organs. The problem is that your scan generation software might produce a scan of a real human. And if you start using that without the person’s permission, you will be in gross violation of their privacy.
Another challenge associated with the training data is that if you want to build an algorithm that works with rare diseases, there will not be much data to work with in the first place.
-
The confidence level must be very high. When human life depends on the algorithm’s performance, there is just too much at stake, which does not leave room for error.
-
Explainability issue. Doctors will not be open to using the ML model if they don’t understand how it produced its recommendations. You can use explainable AI instead, but these algorithms are not as powerful as some black-box unsupervised learning models.
For general AI-associated challenges and implementation tips, check out our article and a free eBook.
To sum up
AI and ML technologies have many applications in the medicine and biology fields. On our blog, you can find more information on artificial intelligence in clinical trials, AI in cancer diagnosing and treatment, and benefits of AI in healthcare.
Bioinformatics is another medicine-related field where ML and AI-based medical solutions come handy. Bioinformatics requires handling large amounts of various data, such as genome sequences, protein structures, and scientific publications. ML is well-known for its data processing capabilities, however, many AI bioinformatics models are expensive to run. It can take hundreds of thousands of dollars to train a deep learning algorithm. For instance, training AlphaFold2 model for protein structure prediction consumes an equivalent of 100-200 GPUs running for several weeks.
You can find more information on what to expect price-wise in our article on how much is costs to implement AI.
If you want to deploy machine learning in bioinformatics, drop us a line. We will work together with you to find the best-suited ML models for a reasonable budget.